|
11/8/2023 0 Comments Palindromic repeats crisprCRT's approach to detecting repetitive sequences is straightforward. This tool, CRT, was shown to be a significant improvement over the current techniques for CRISPR identification. In this paper a new tool was introduced for the automatic detection of CRISPR elements. In terms of speed, CRT also demonstrated superior performance, especially for genomes containing large numbers of repeats. When compared to Pilercr, CRT showed improved performance for recall and quality. In terms of correctness, CRT was shown to be very reliable, demonstrating significant improvements over Patscan for measures precision, recall and quality. CRT was compared to CRISPR detection tools, Patscan and more » Pilercr. In this study, a new tool, CRT, is introduced that rapidly and accurately identifies CRISPRs in large DNA strings, such as genomes and metagenomes. Existing repeat detection tools do a poor job of identifying CRISPRs due to the presence of unique spacer sequences separating the repeats. CRISPRs are beginning to attract attention because of their proposed mechanism that is, defending their hosts against invading extrachromosomal elements such as viruses. « lessĬlustered Regularly Interspaced Palindromic Repeats (CRISPRs) are a novel type of direct repeat found in a wide range of bacteria and archaea. Thus, CAS2 proteins are sequence-specific endoribonucleases, and we propose that their role in the CRISPR-mediated anti-phage defense might involve degradation of phage or cellular mRNAs. 
Mutagenesis of SSO1404 identified six residues (Tyr-9, Asp-10, Arg-17, Arg-19, Arg-31, and Phe-37) that are important for enzymatic activity and suggested that Asp-10 might be the principal catalytic residue. Here we characterize the Enterococcus faecalis Csn2 protein as a double-stranded (ds-) DNA-binding protein and report its 2.7 resolution revealing the first ribonuclease with a ferredoxin-like fold. 
Csn2 is a Nmeni subtype-specific cas gene required for new spacer acquisition. A key molecular event during this process is the acquisition of new spacers into the CRISPR loci to guide the selective degradation of the matching foreign genetic elements. 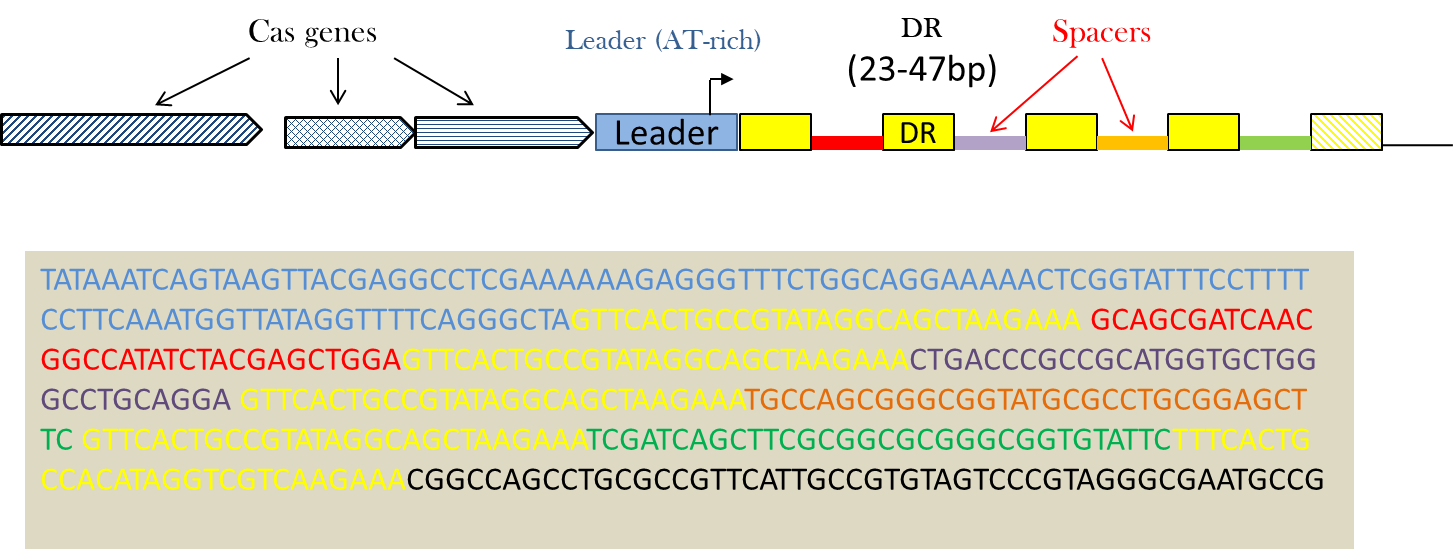
They form a line of RNA-based immunity to eradicate invading bacteriophages and malicious plasmids. Clustered regularly interspaced short palindromic repeats (CRISPR) and their associated protein genes (cas genes) are widespread in bacteria and archaea.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
